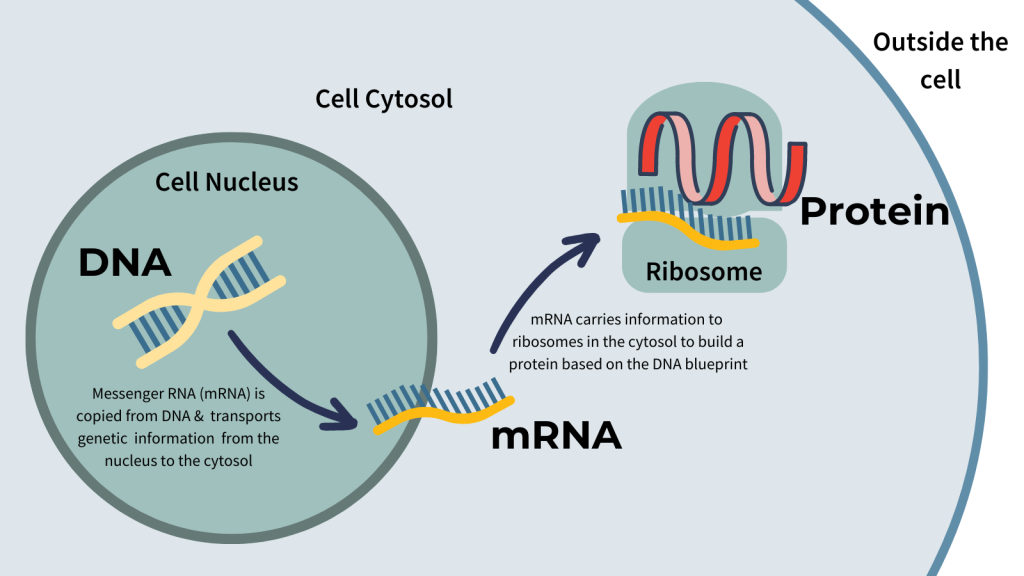
Green fluorescent protein (GFP) has become a widely used direct detection marker in live cell experiments. It is a protein that fluoresces brightly upon excitation at 491 nm and emits light maximally at 503 nm.
mRNA electroporation rapidly increased the protein expression level, which peaked within 1 h after treatment. The results correlated well with the DegScore values, which predict mRNA stability.
Molecular Structure
The EGFP gene encodes an enhanced version of the green fluorescent protein (GFP) from the jellyfish Aequorea victoria. eGFP is an extraordinarily bright and stable GFP variant that can be used in mammalian cells as a direct detection reporter.
eGFP has a wide spectral range with a maximum excitation wavelength of 491 nm and a stokes shift of 503 nm. Unlike wt GFP, which had many drawbacks that made it unsuitable for cell imaging, eGFP is highly stable, pH-insensitive, and aggregation-resistant.
To improve the stability of eGFP, an mRNA is created with a twin ribozyme-based repair system that replaces single nucleotides in a full-length mRNA transcript with an externally added patch. This system was shown to restore a functional EGFP protein, as measured by restored fluorescence.
The crystal structure of EGFP at 1.35 A resolution showed that the overall structures are similar to those of wt GFP, with a classic b-barrel fold and a chromophore helix running through the core. The F64L and S65T mutations have a significant impact on the hydrogen bond network surrounding the chromophore, with the replacement of Phe64 leading to subtle rearrangements of hydrophobic core packing that reduce surface exposure of two critical residues in the chromophore charge defining the position of Glu222.
RNA Structure
The chromophore of GFP is tightly packed within the core of the b-barrel structure, protecting it from quenching through water dipoles, paramagnetic oxygen, or cis-trans isomerization. This packing is further tuned through non-covalent interactions with neighbouring residues. The F64L mutation alters these interactions by swapping the bulky and buried phenylalanine for a leucine residue in the central chromophore-containing helix. This allows the helix to pack closer with the chromophore but results in a polar shift in the position of residue 222. The buried side chain of Glu222 is positioned in two different conformers; the position of the carboxylate group in conformer A is superimposable to that observed in wt GFP, and the recently determined lower resolution structure of a His-tagged S65T GFP mutant 2Y0G. The other conformation, however, is significantly different concerning wt GFP and results in a new side chain positioning for the molecule.
The higher B-factor values for EGFP 4EUL relative to those observed in the lower resolution structures of both wt GFP and the S65T GFP mutant result in increased confidence in the placement of side chains during model building. Interestingly, the Thr65 hydroxyl group in EGFP is positioned slightly differently than that of wt GFP, but this can be explained by steric effects on the position of the carboxylate group caused by adding an extra methyl group to Glu.
mRNA Function
EGFP is an enhanced version of GFP (green fluorescent protein), originally isolated from jellyfish Aequorea Victoria, and shows bright green fluorescence with an emission peak at 509 nm. It is a widely used direct detection reporter for monitoring protein expression and localization in mammalian cells. EGFP mRNA is capped with a proprietary co-transcriptional capping method and polyadenylated to mimic fully processed mature mRNA. Vernal Biosciences eGFP mRNA is optimized for use in various functional studies.
This mRNA can drive the expression of the enhanced green fluorescent protein (EGFP) in cells. The mRNA is delivered using lipid nanoparticles (LNPs) composed of SM-102, DSPC, cholesterol, and DMG-PEG2000 at optimal molar concentration for rapid encapsulation and efficient mRNA delivery.
The LNPs are then infused into cells by either electroporation or transfection. The mRNA is translated into the corresponding protein, which can be monitored through ratiometric fluorescence measurements using a luminometer. Fluorescence decay data correlated well with measured in-solution half-life and moderately with 24-h protein expression. However, 6-h protein expression exhibited a less clear correlation with these data. This may be due to other factors impacting mRNA translation and stability, such as endogenous proteins inhibiting autophagic fusion.
mRNA Degradation
Integrating mRNA transcription and decay processes is critical to shaping appropriate gene expression patterns during cellular proliferation, differentiation, and stress responses. This process ensures that a limited number of transcripts are available for translation at any given time, thereby controlling the cell’s rate at which proteins are produced.
Consequently, the dynamics of mRNA degradation directly determine how swiftly changes in transcriptional rates are reflected at the protein level. Long mRNA transcripts with fast turnover will quickly attenuate the effect of changes in transcriptional rates, whereas short mRNA transcripts with slow turnover will be less affected.
It is, therefore, crucial to understand the mechanisms underlying mRNA degradation. Several studies have identified sequence motifs that correlate with mRNA stability, including the AU-rich element (ARE), which has been shown to promote mRNA degradation.
Moreover, other factors such as promoters and transcriptional efficiency also influence the decay rate of mRNAs. The addition of glucose in the yeast strain Saccharomyces cerevisiae triggers a massive and selective transcriptional burst that is immediately attenuated by a mRNA decay mechanism involving the native GAL genes.
In general, most mRNAs are degraded by a deadenylation-dependent process. The mRNA’s poly(A) tail is initially shortened by the nine-subunit Ccr4p/Pop2/Not complex and then decapped by the Dcp1p/Pat1p/Lsm complex. mRNAs with a functional 3′-5′ exonuclease, such as Xrn1p, will also undergo a secondary decapping reaction before being targeted for degradation by the cytoplasmic exosome and Ski7p.